The Pool lab uses genomic diversity to learn about natural selection and the histories of populations. One major focus of our research is to understand how adaptive evolution operates at the genetic level. We do this using our extensive collection of wild-derived Drosophila melanogaster strains, comparing populations that are closely related but have adapted to contrasting environments. We also have a strong focus on population genomic analysis and methods, and we have a growing interest in leveraging D. melanogaster to study the genetic basis of partial reproductive isolation. Here are some basic explanations about why flies are an ideal study system for our research and why population genetics is such a unique field of study, while below you can read more about our ongoing projects.
How does adaptive evolution operate at the genetic level in nature?
~ When a trait evolves, how many genes contribute?
~ Do the causative variants tend to reach 100% frequency?
~ Does natural selection usually act on new mutations or standing genetic variation?
~ Is adaptive evolution genetically predictable?
The Pool lab has a special emphasis on local adaptation (traits and underlying alleles that have a geographically limited advantage). Local adaptation gives us the clearest possible window into how adaptive evolution operates in nature: we can use crosses to map the causative genomic regions, we can analyze genetic variation to find differentially selected genes, and the unrivaled molecular tool kit of D. melanogaster can help to confirm and characterize the causative genes and variants. We follow just this pipeline to investigate the genetic underpinnings of a wide range of locally adaptive traits, including melanic pigmentation, large body and wing size, and elevated cold and ethanol tolerance. An emerging portrait from our studies is that selection may frequently act on standing genetic variants, including many of moderate to large effect, but the genetic architecture of these adaptive traits often remains variable within the evolved populations (Bastide et al. 2016; Sprengelmeyer & Pool 2021; Sprengelmeyer et al. 2022; unpublished data). These studies have often involved a fusion of quantitative trait locus (QTL) mapping with population genetic scans for local adaptation. Now, we are leveraging the persistent genetic variability underlying by these adaptive traits by using genotype-phenotype association testing to narrowly localize the causative genetic variants.
Building on our past work on the contributions of gene expression levels and alternative splicing to the (parallel) evolution of cold tolerance within
D. melanogaster (Huang et al. 2021; 2022), we are now conducting a broader evolutionary "multi-omic" study to discover which level of gene regulation are the most responsive to adaptive evolution.
To complement our experimental efforts, we also maintain a strong focus developing new statistical methods and computational resources to extract new insights from genomic data sets. Our projects in this realm have included (1) developing the Genome Nexus (Lack et al. 2015; 2016) (2) new statistics for detecting local adaptation from genetic variation (Lange et al. 2016; Yassin et al. 2016; da Silva Ribeiro et al. 2022), and (3) a new method for analyzing bulk mapping data (Pool 2016),
Of course, gaining a strong understanding of how natural selection has acted at specific genes within a species relies on a clear view of how population history has shaped its genome-wide genetic diversity. The lab has played a leading role in uncovering this history of this key model species, including by analyzing the genomes of newly-discovered wilderness-living flies from the species' African ancestral range, and by estimating a detailed historical model of the geographic expansion of human-associated populations (Sprengelmeyer et al. 2020). We have also zoomed in on key recent time intervals by analyzing genomes from museum-preserved flies (Shpak et al. 2023).
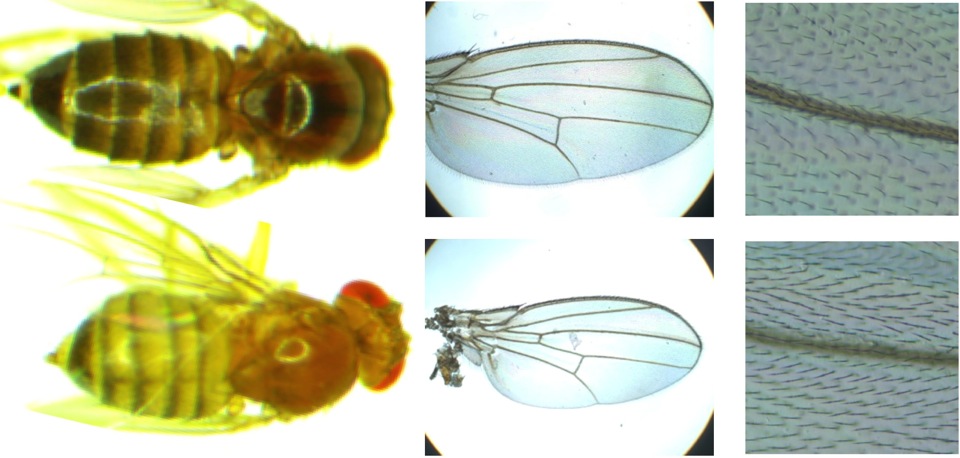

How does one species begin to become two?
~ How quickly does reproductive isolation begin to develop between populations?
~ How important is natural selection in mediating admixture between populations?
~ Which genes and biological processes contribute to incompatibilities between populations?
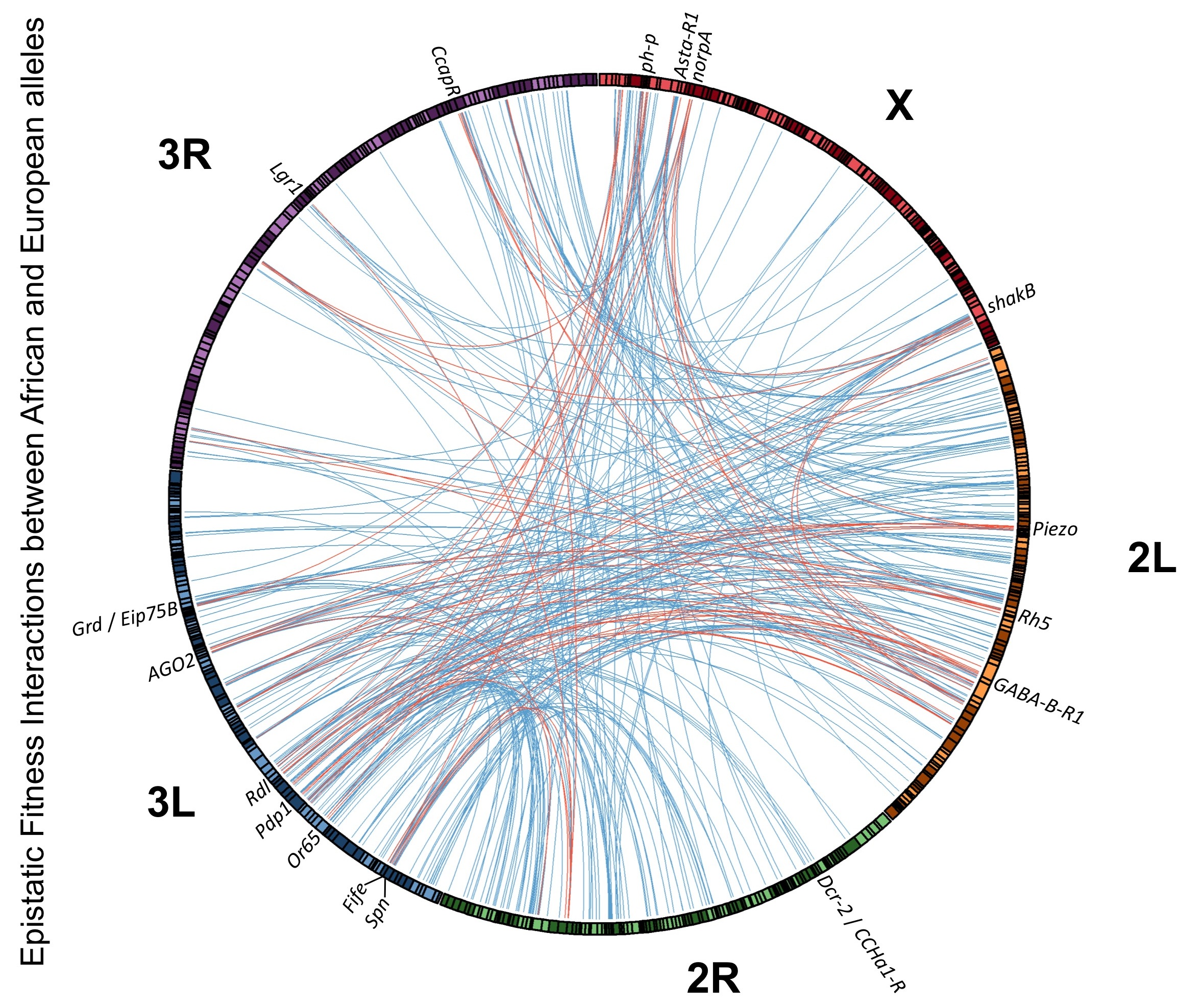
While reproductive isolation is often studied betwen taxa that are almost completely unable to interbreed, there could be considerable advantages to studying how the first steps toward reproductive isolation occur. Perhaps surprisingly, the fly populations we study from places like Africa vs. Europe, although they shared a common ancestor just ~13,000 years ago, have already begun to show signs of reproductive isolation in the form of "hybrid breakdown" likely driven by genetic incompatibilities. Our interest in this topic sprang up accidentally, upon showing that the North Carolina "DGRP" population (which has a blend of European and African ancestry) shows patterns of genomic ancestry strongly influenced by natural selection, including a genome-wide abundance of "ancestry disequilibrium" between unlinked loci that could reflect numerous incompatibilities (Pool 2015). We subsequently confirmed one form of hybrid breakdown experimentally, finding that some Africa x Europe crosses yielded relatively high numbers of sterile F2 males (Lollar et al. 2023) that may involve complex incompatibilities involving uniparentally-inherited genetic factors. This work sets the stage for our ongoing research mapping the genetic loci underlying the incompatibilities between African and European alleles that lead to male sterility, and illuminating the biological basis of this instance of hybrid breakdown.